1. Long Non-coding RNA Identification
Click button “Run” to enter the page of lncRNA identification. This page contains two parts: run web server to identify lncRNAs and download previous results.
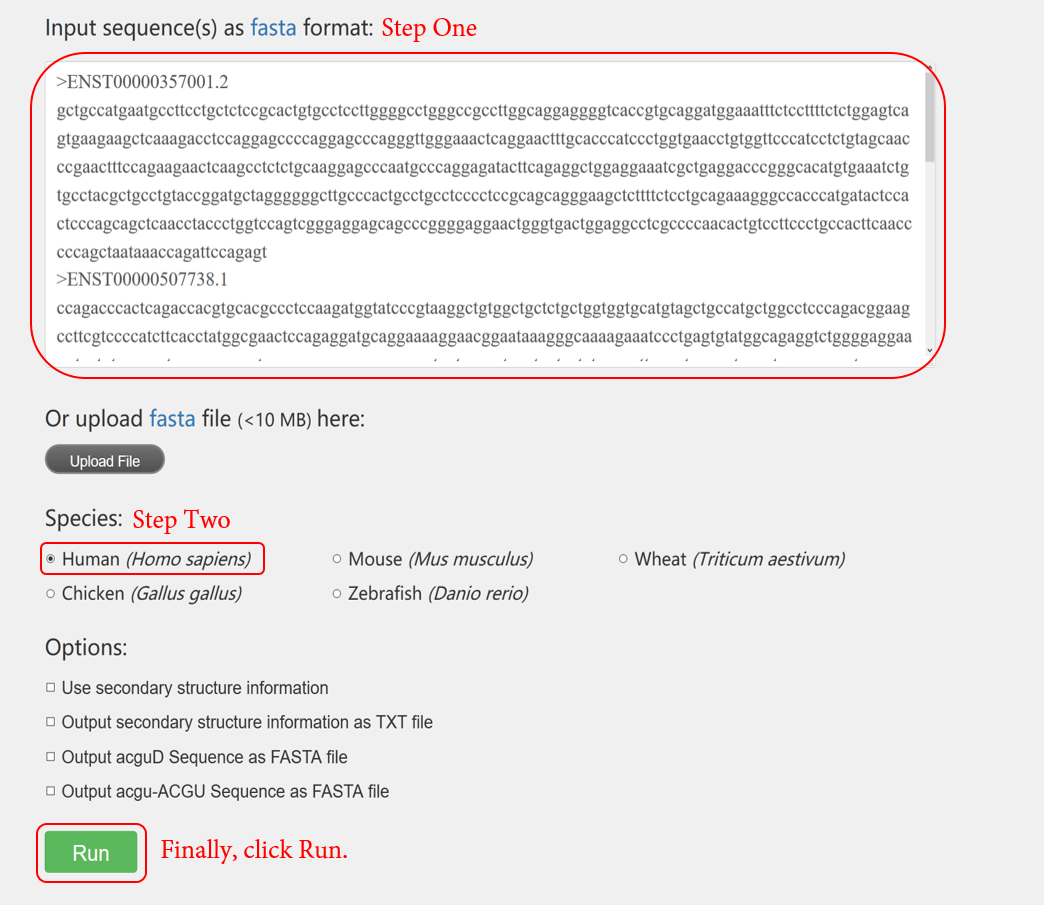
1.1.1 In this example, we choose to input sequences in text area. The input format need to be in FASTA format. Identify sequence using sequence and EIIP-derived features only.
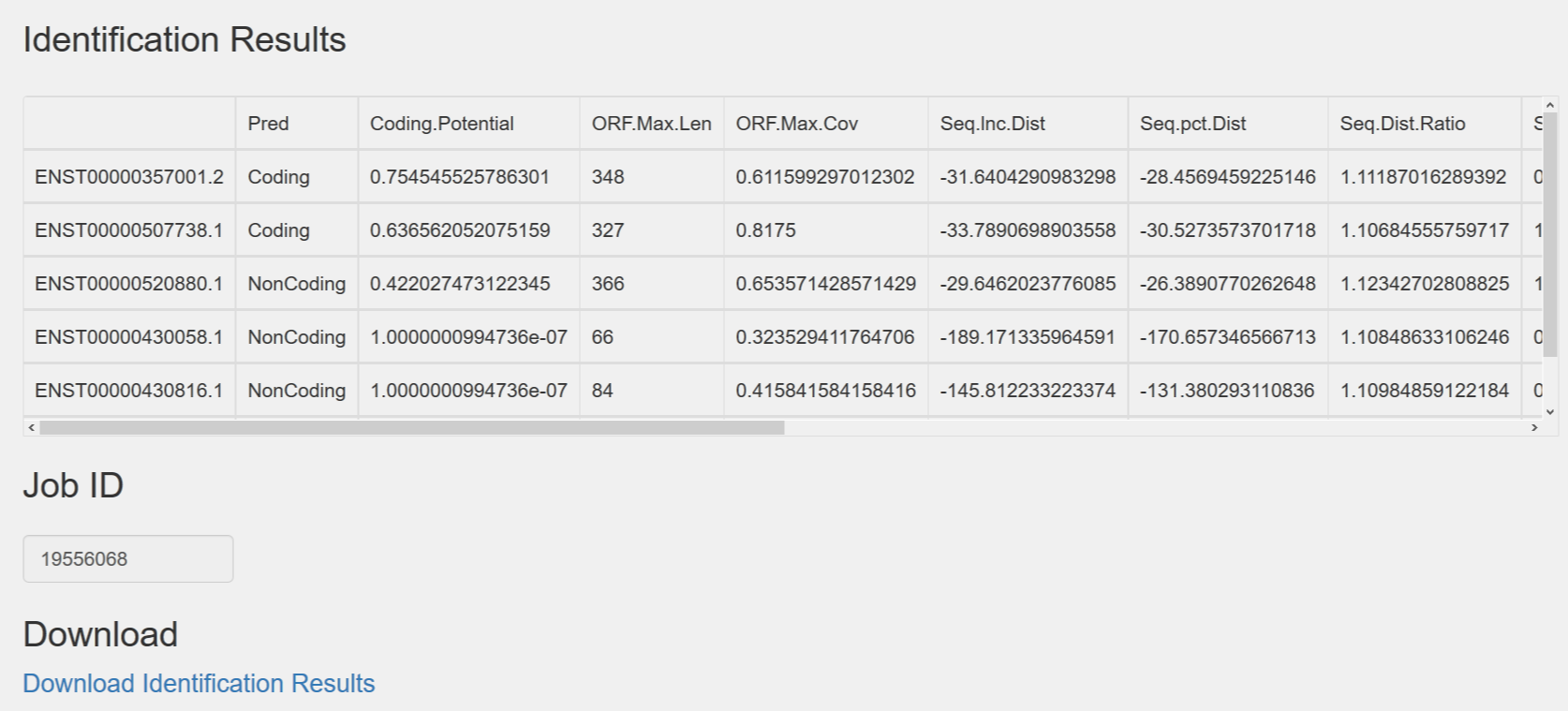
1.1.2 The values of different features are presented in the table. Job ID can be used to re-download result files. The original result data are saved as *.csv file which can be analysed easily in multiple software.
1.2.1 In this example, we firstly upload a FASTA file and predict the sequences with all available features. Here, we select species “mouse” and predict lncRNAs with secondary structure-derived features. We check all the options to output secondary structure information as well. The time cost is mainly determined by “RNAfold”. The process of calculating secondary structure of long transcripts may be time-consuming, but here only the sequences with length < 4000 nt will be computed. This parameter can be customized using LncFinder's R package.
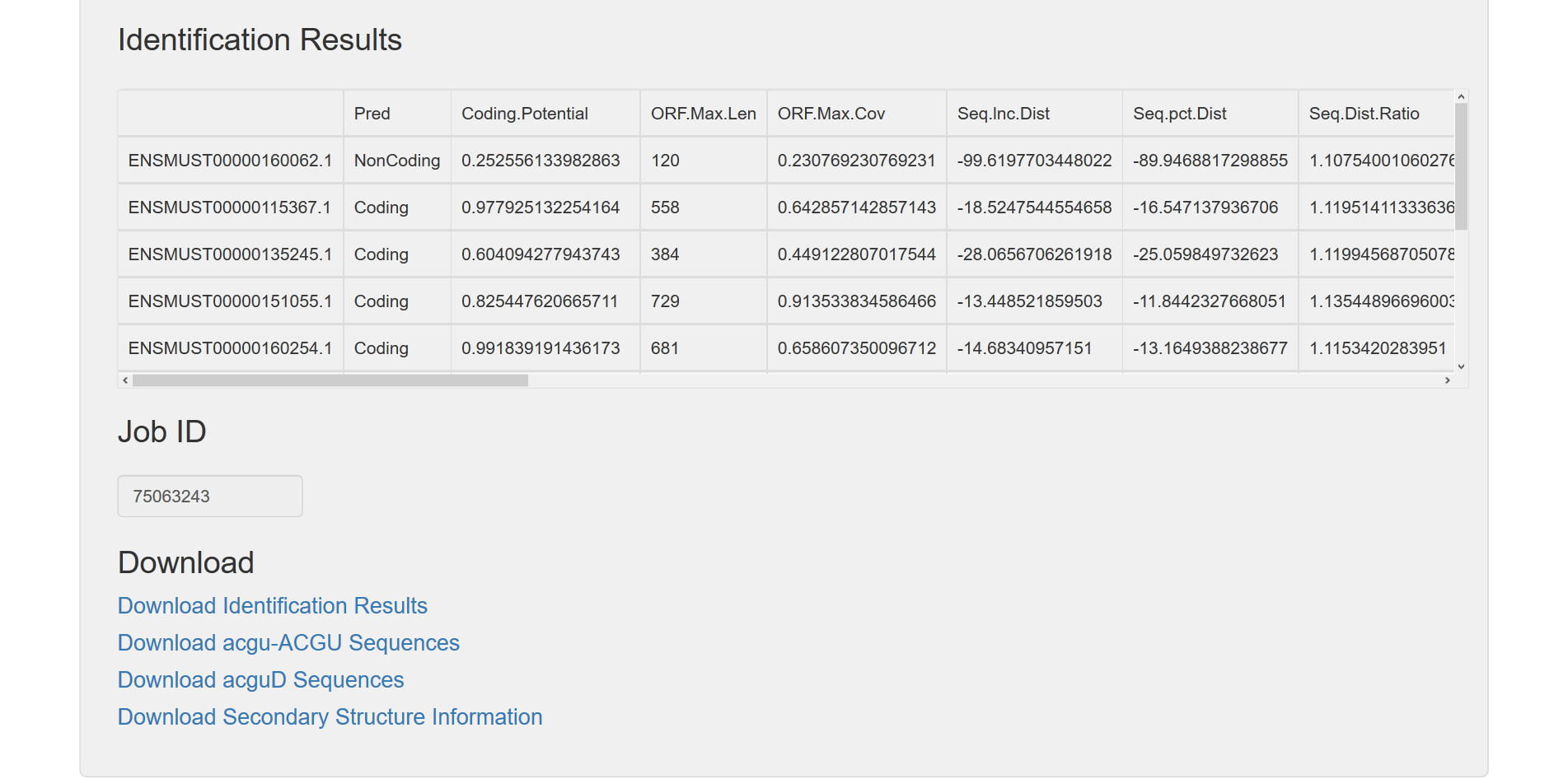
1.2.2 Download links are provided based on the checked options. Here, result files: secondary structure information, acguD Seq and acgu-ACGU Seq are available.
2. Download previous results
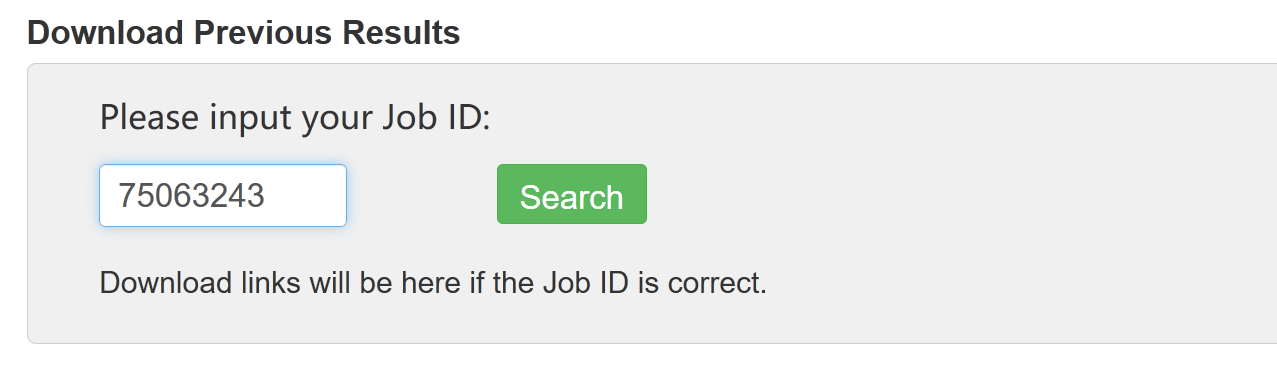
Users can use Job ID to download previous result data:
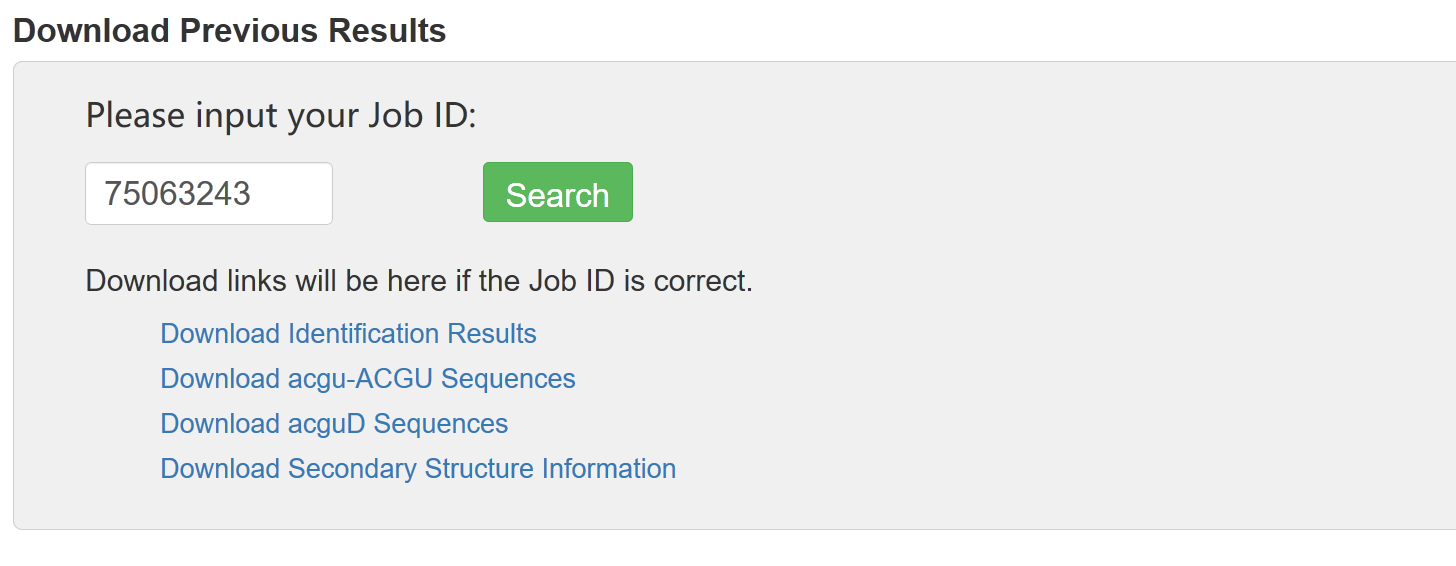
And the related files will be displayed at the same page:
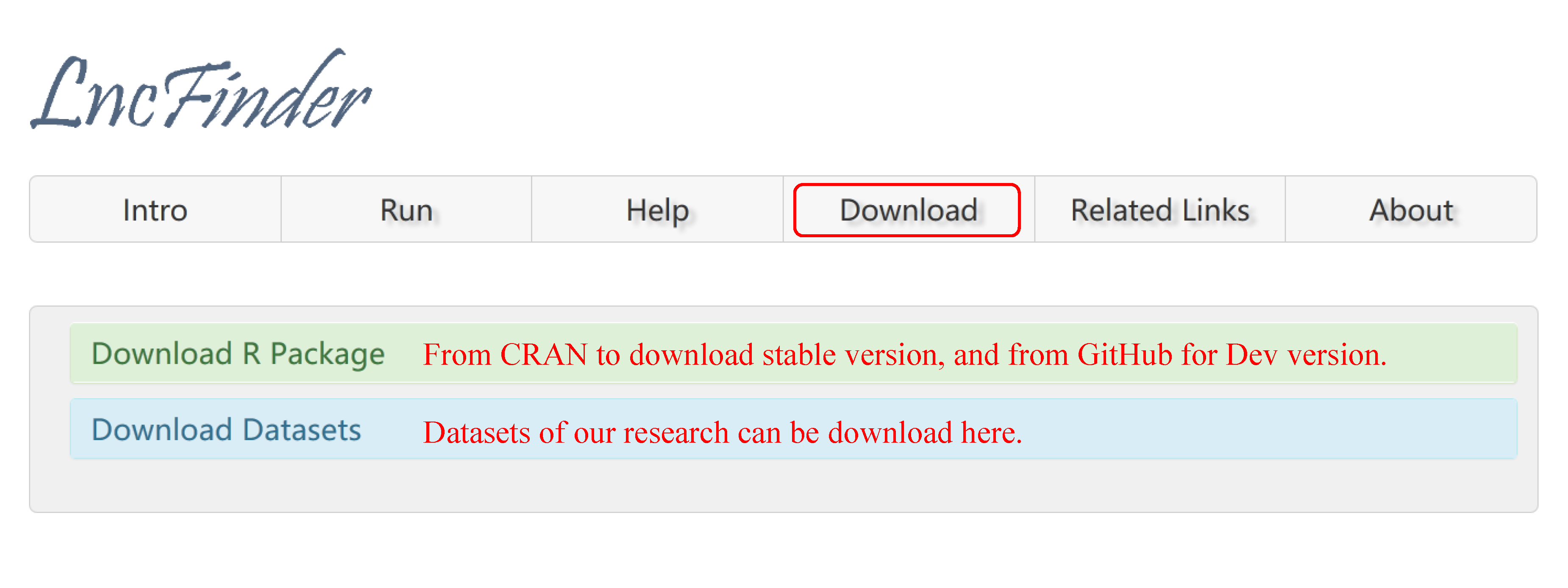
3. Download of R package and Datasets
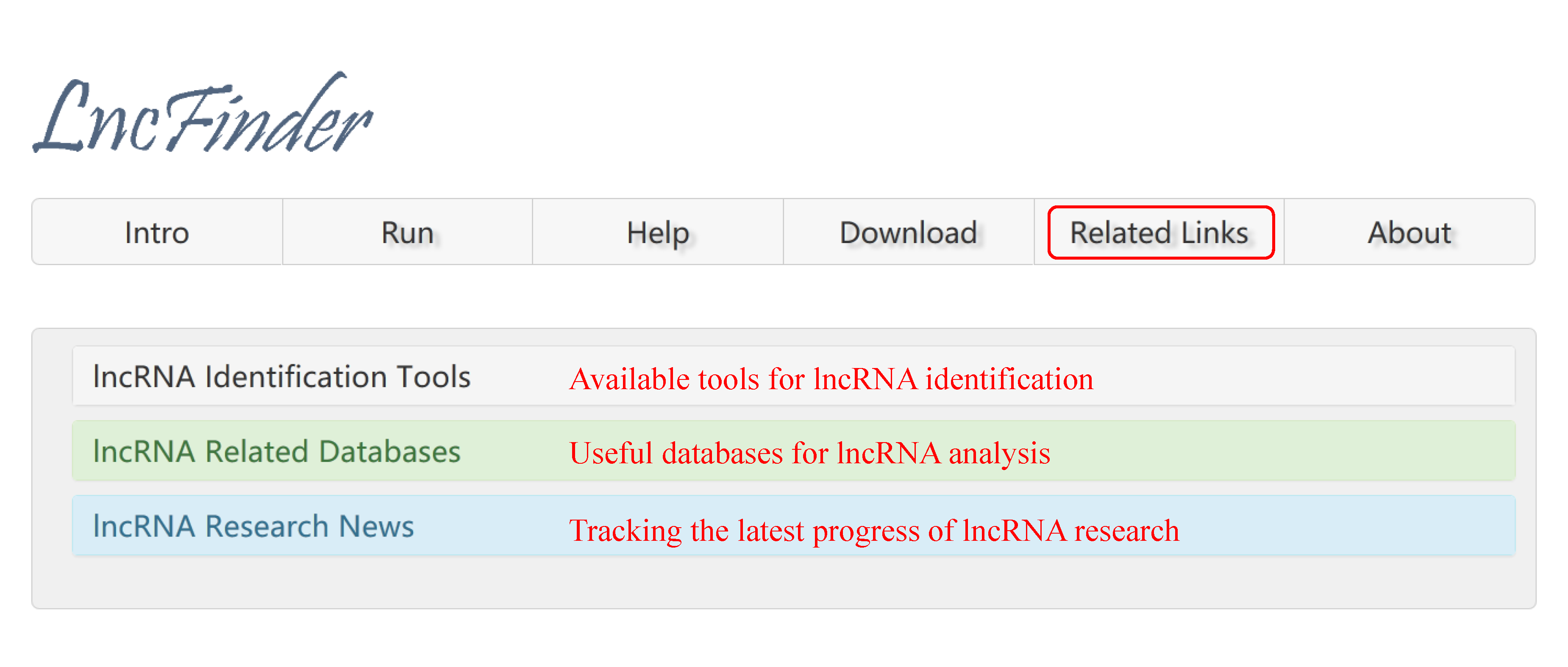
4. LncRNA-related Links